library(MASS)
data(cats)
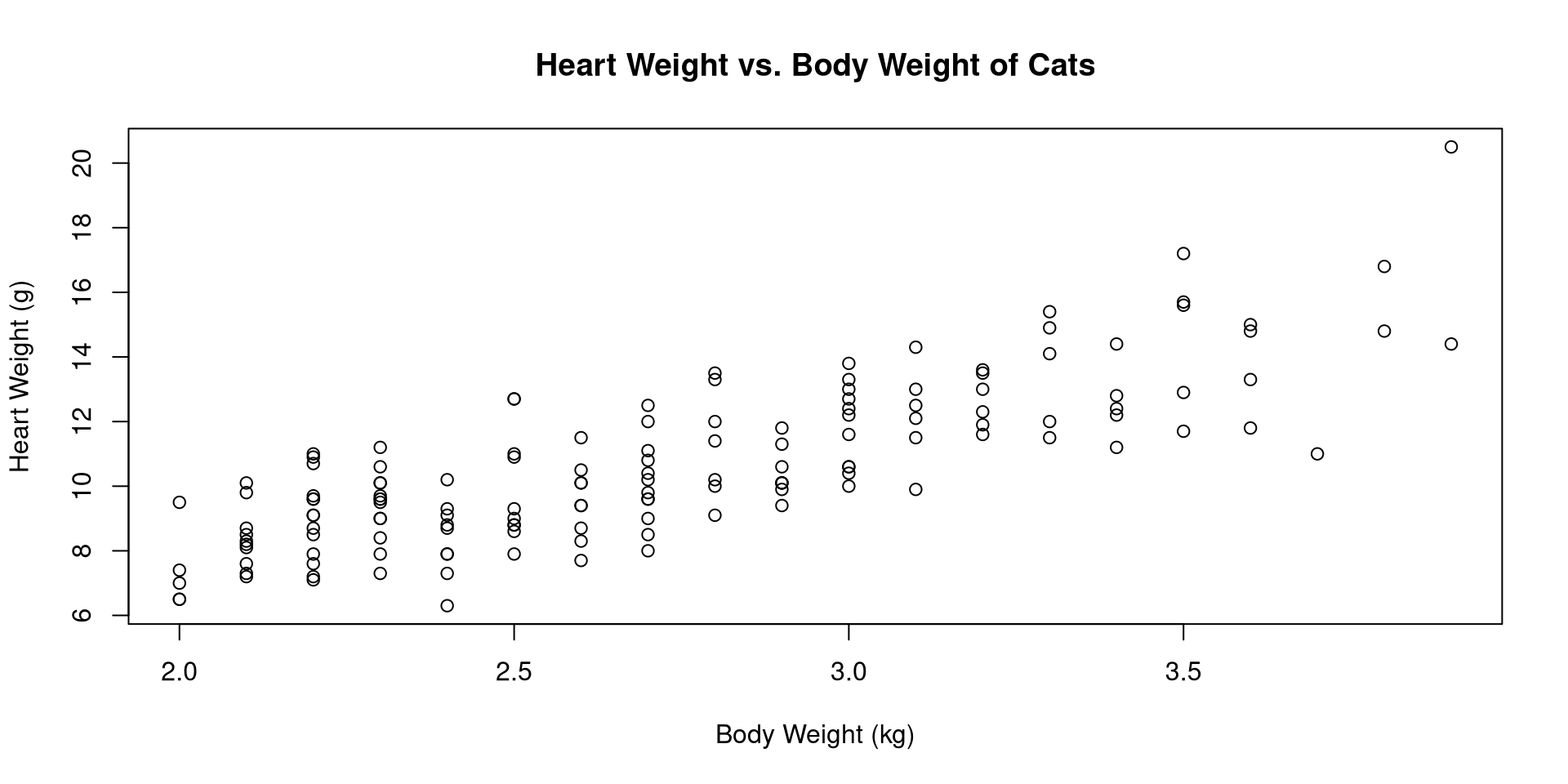
with(cats, plot(Bwt, Hwt, xlab="Body Weight (kg)",
ylab="Heart Weight (g)",
main="Heart Weight vs. Body Weight of Cats"))
Pearson's product-moment correlation
data: Bwt and Hwt
t = 16.119, df = 142, p-value < 2.2e-16
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
0.7375682 0.8552122
sample estimates:
cor
0.8041274
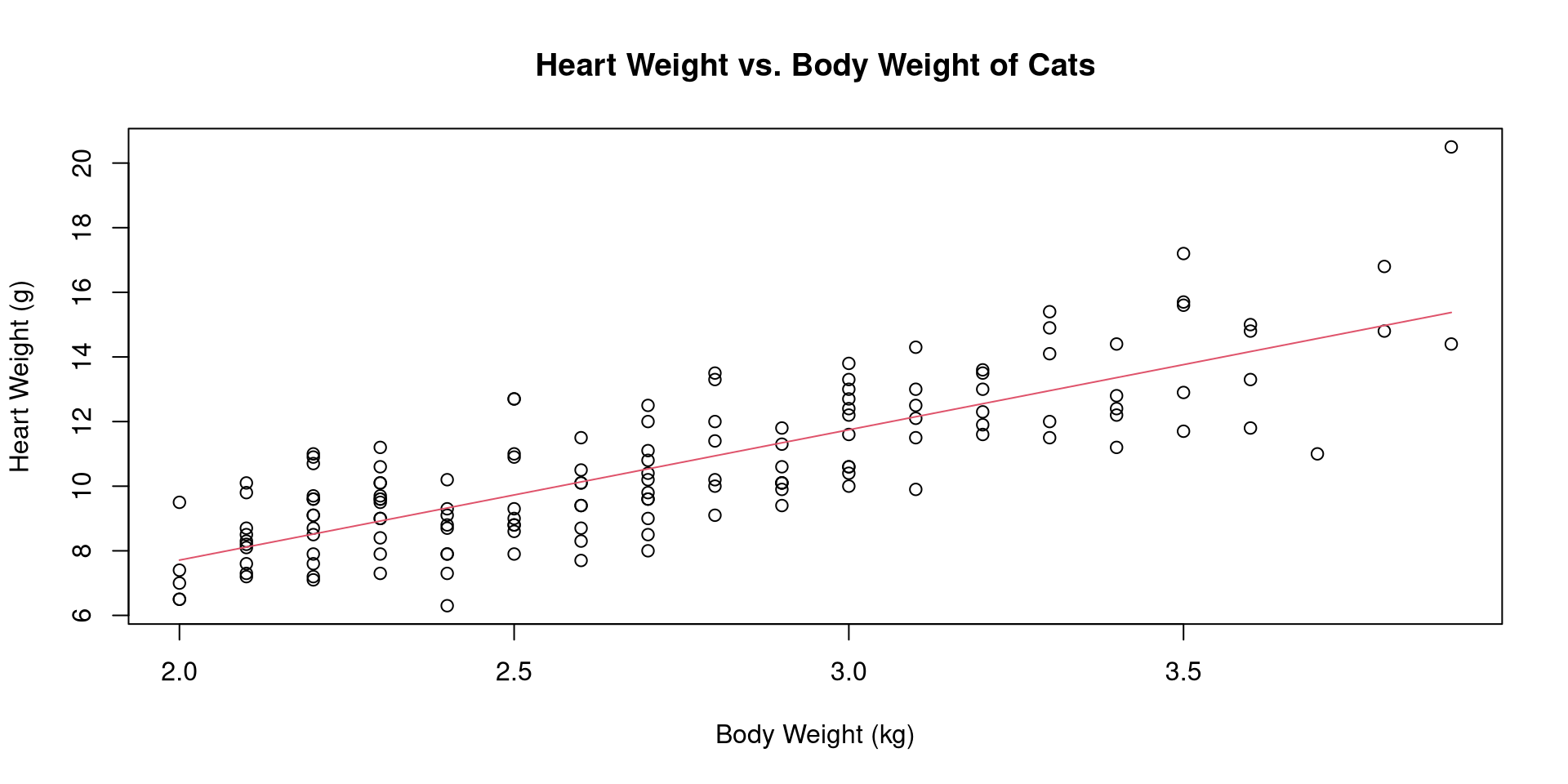
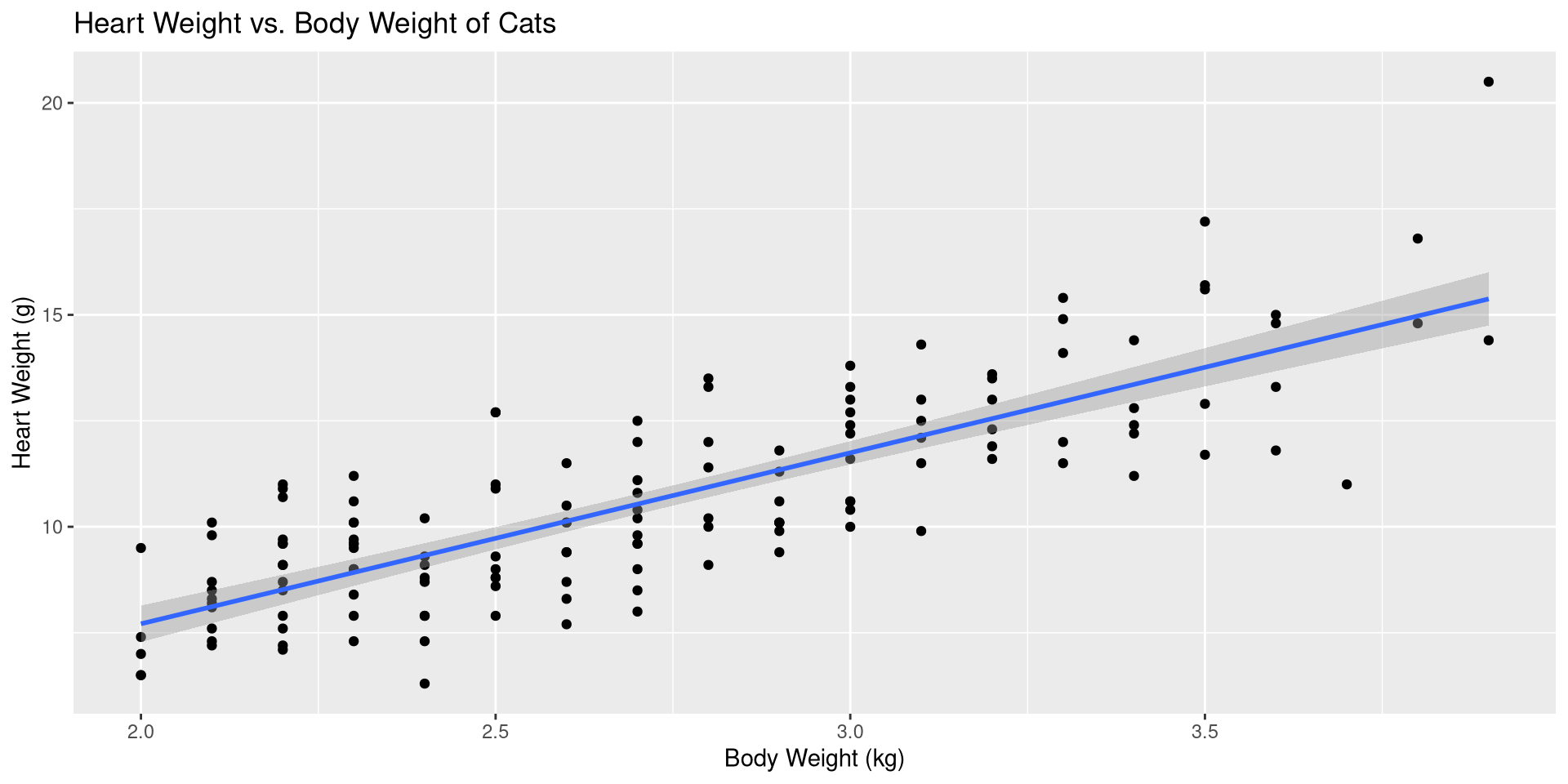
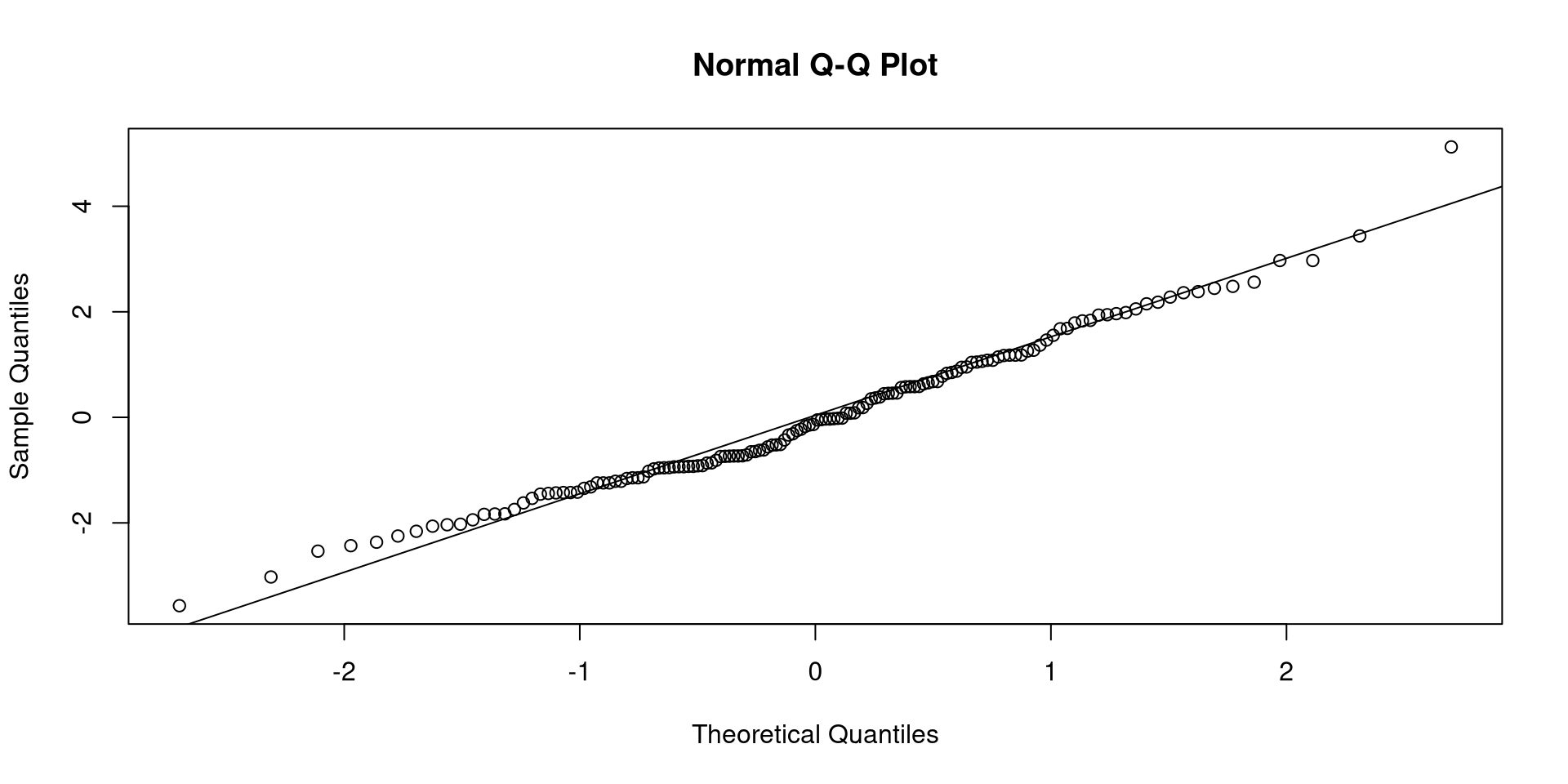
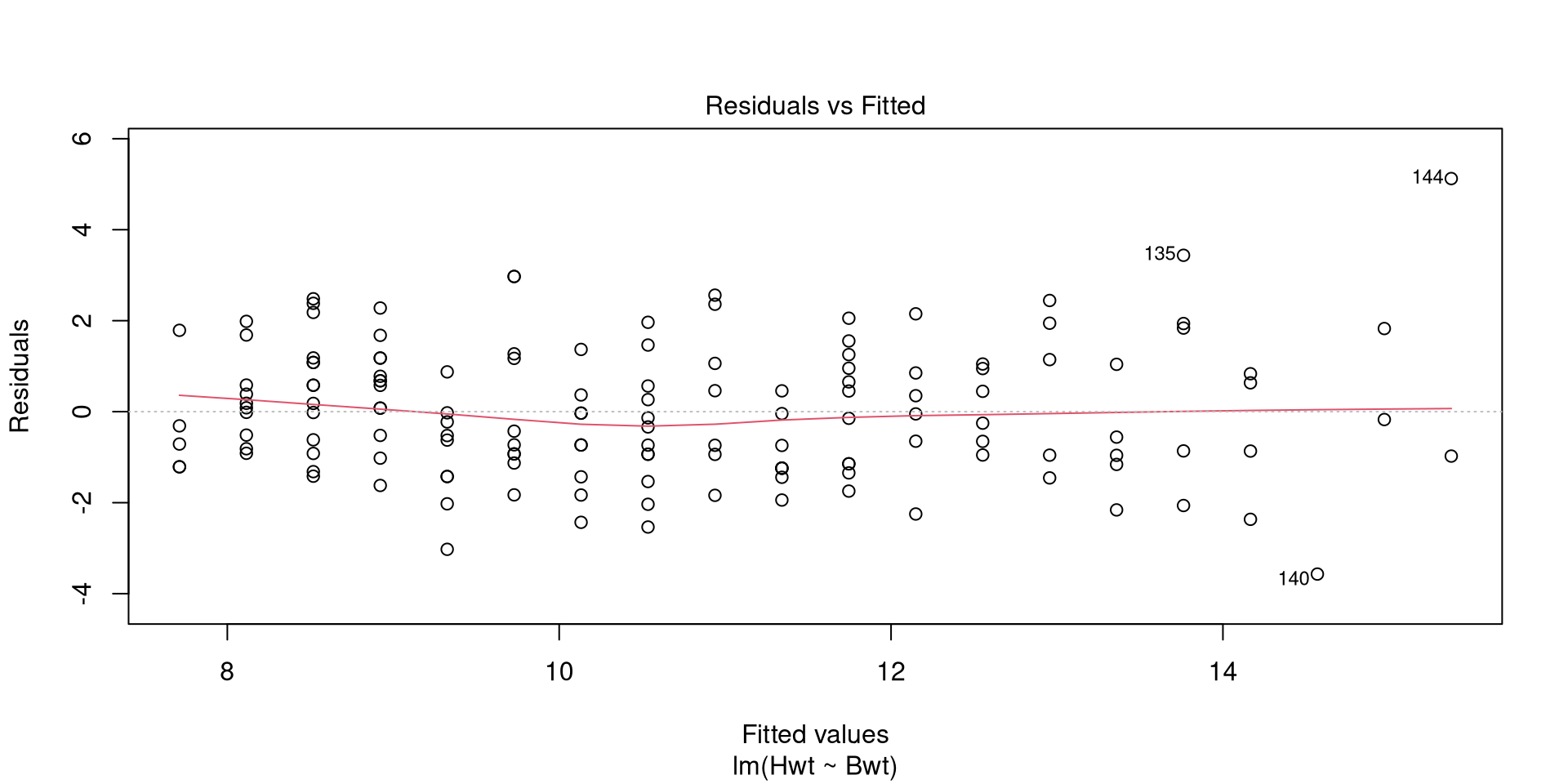
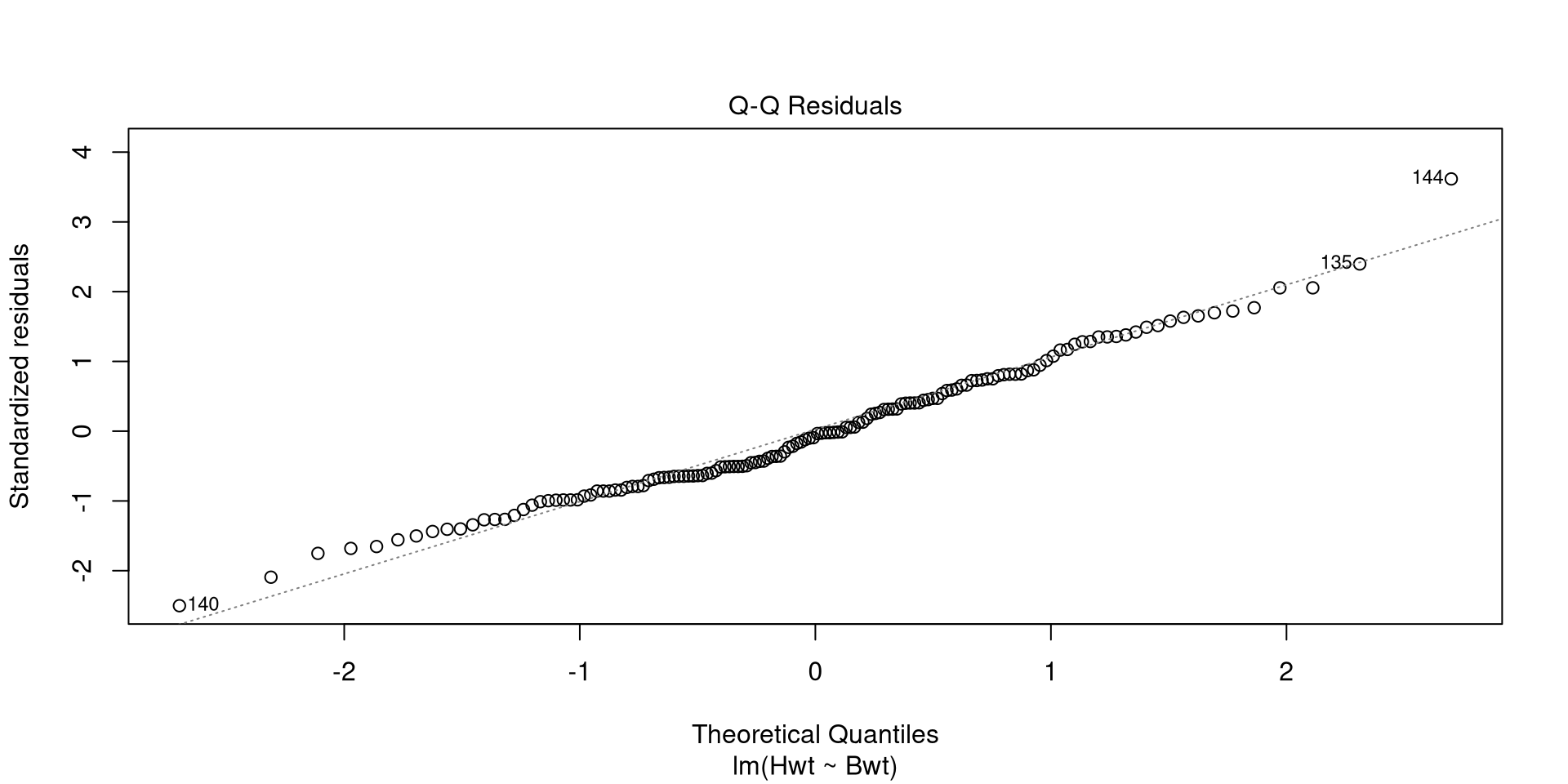
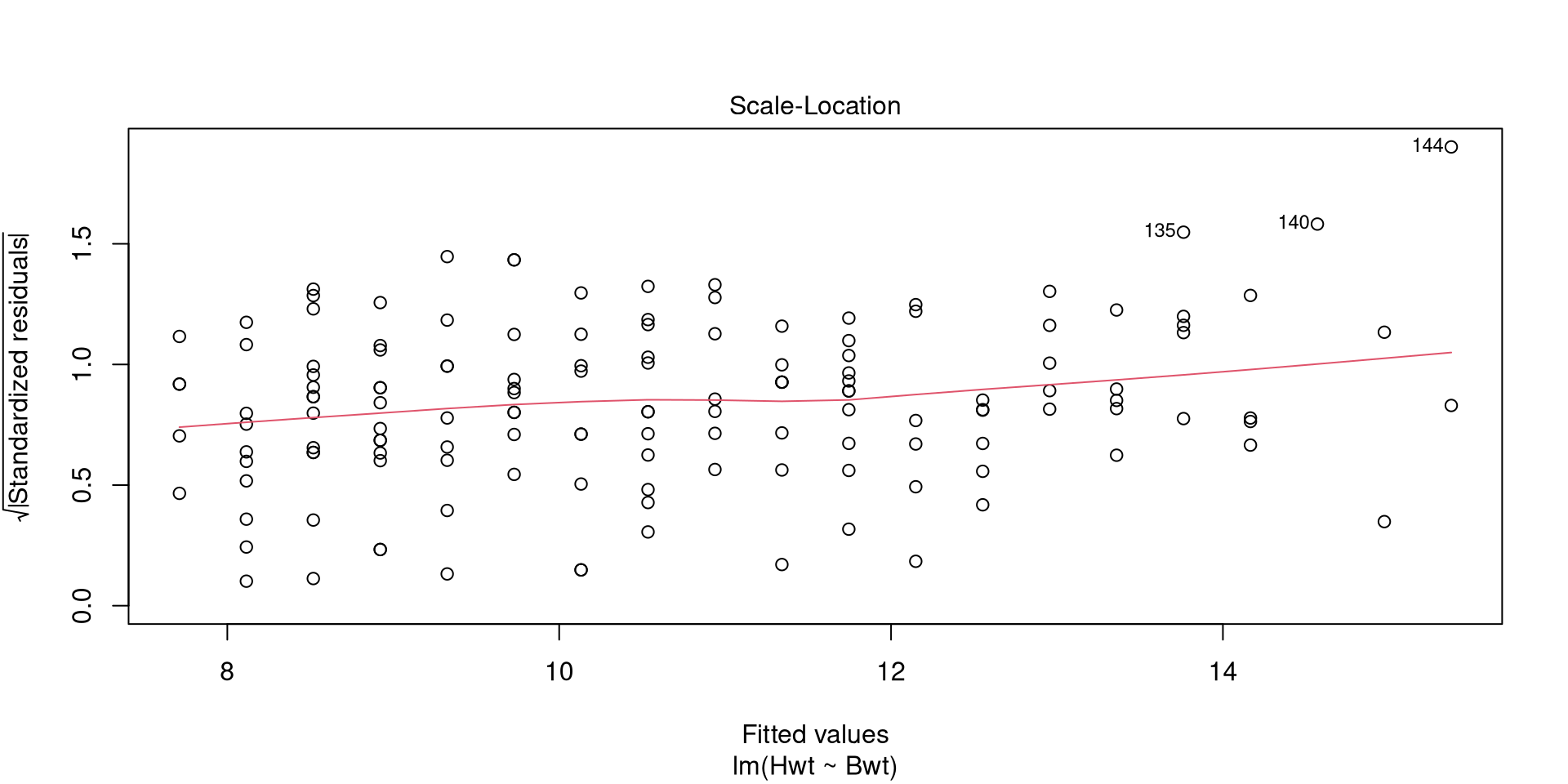
Call:
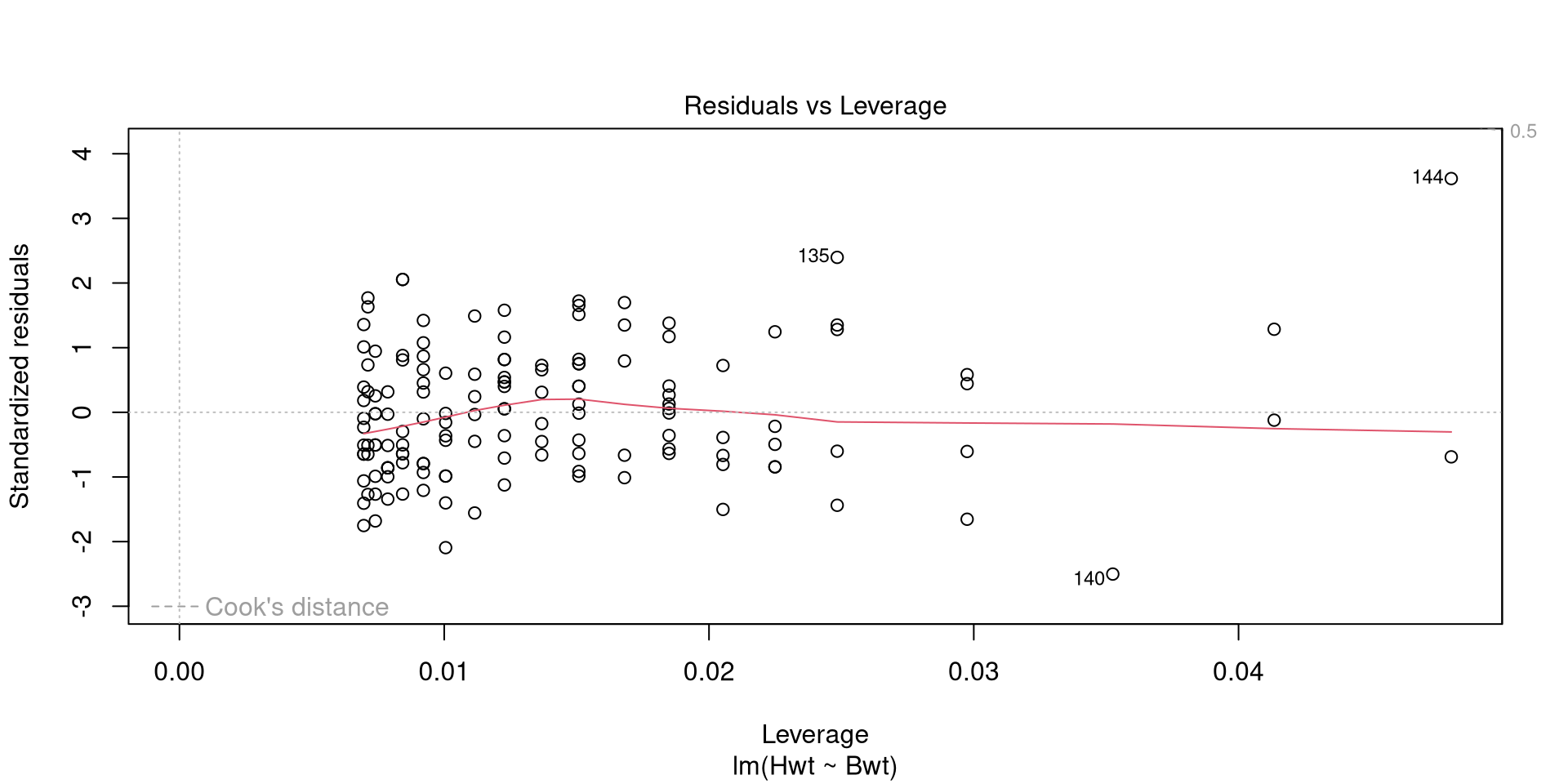
lm(formula = Hwt ~ Bwt, data = cats)
Coefficients:
(Intercept) Bwt
-0.3567 4.0341