dat <- read.csv("NHANES1.csv")
# make factors
integer_info <- sapply(dat, is.integer)
integer_info[which(names(integer_info) == "age")] <- FALSE # age should stay an integer
dat[integer_info] <- lapply(dat[integer_info], as.factor)Exercise 4: Solutions 5th part
Load the data in R.
Task 4.1
Use a chi-square test in order to test whether the presence of chronic bronchitis and the current smoking status are independent.
table(dat$cbronch_now, dat$smokstat)
1 2 3
FALSE 37 8 28
TRUE 32 3 43chisq.test(dat$cbronch_now, dat$smokstat)
Pearson's Chi-squared test
data: dat$cbronch_now and dat$smokstat
X-squared = 5.6447, df = 2, p-value = 0.05947Task 4.2
Use a Fisher test to verify the independence between sex and the presence of any kind of liver disease.
table(dat$livdis_now, dat$male)
FALSE TRUE
FALSE 33 38
TRUE 50 60fisher.test(dat$livdis_now, dat$male)
Fisher's Exact Test for Count Data
data: dat$livdis_now and dat$male
p-value = 1
alternative hypothesis: true odds ratio is not equal to 1
95 percent confidence interval:
0.547707 1.979023
sample estimates:
odds ratio
1.041893 Task 4.3
Perform a sign test both on hdl and on log-hdl to test the hypothesis that the median of the cholesterol level is 1.30. Is the median significantly different from 1.30? Do you obtain the same results using hdl and logHdl?
library(BSDA)Loading required package: lattice
Attaching package: 'BSDA'The following object is masked from 'package:datasets':
OrangeSIGN.test(dat$hdl, md = 1.30)
One-sample Sign-Test
data: dat$hdl
s = 2180, p-value = 0.3286
alternative hypothesis: true median is not equal to 1.3
95 percent confidence interval:
1.29 1.32
sample estimates:
median of x
1.29
Achieved and Interpolated Confidence Intervals:
Conf.Level L.E.pt U.E.pt
Lower Achieved CI 0.9475 1.29 1.32
Interpolated CI 0.9500 1.29 1.32
Upper Achieved CI 0.9511 1.29 1.32SIGN.test(log(dat$hdl), md = log(1.30))
One-sample Sign-Test
data: log(dat$hdl)
s = 2180, p-value = 0.3286
alternative hypothesis: true median is not equal to 0.2623643
95 percent confidence interval:
0.2546422 0.2776317
sample estimates:
median of x
0.2546422
Achieved and Interpolated Confidence Intervals:
Conf.Level L.E.pt U.E.pt
Lower Achieved CI 0.9475 0.2546 0.2776
Interpolated CI 0.9500 0.2546 0.2776
Upper Achieved CI 0.9511 0.2546 0.2776Task 4.4
Use a Mann-Whitney test to test the null hypothesis \(H_0 : male weight = female weight\).
library(coin)Loading required package: survivalwilcox_test(weight ~ as.factor(male), data = dat)
Asymptotic Wilcoxon-Mann-Whitney Test
data: weight by as.factor(male) (FALSE, TRUE)
Z = -18.398, p-value < 2.2e-16
alternative hypothesis: true mu is not equal to 0Task 4.5
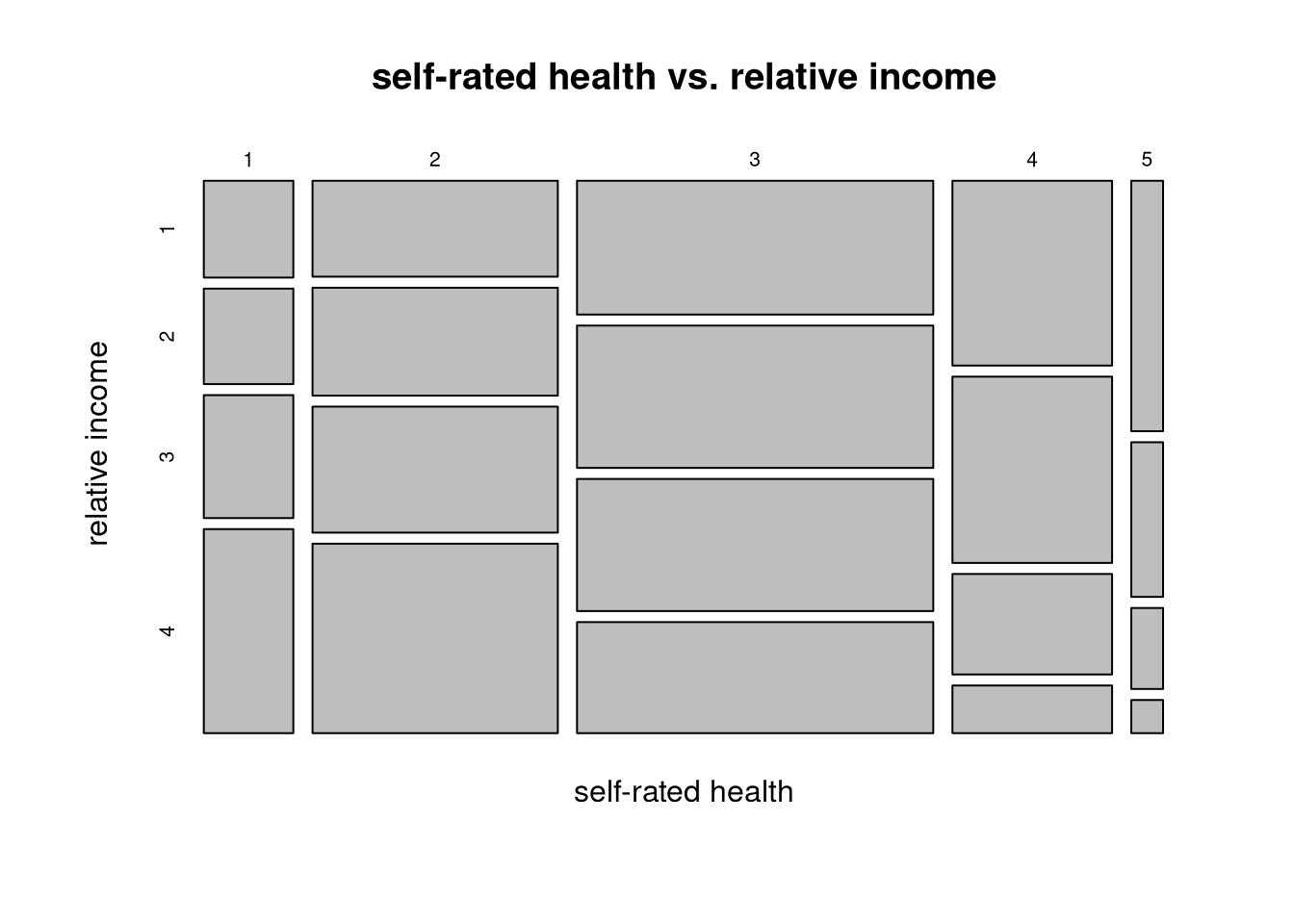
Task 4.6
Test the relation for statistical significance using an appropriate test.
chisq.test(dat$srhgnrl, dat$increl)
Pearson's Chi-squared test
data: dat$srhgnrl and dat$increl
X-squared = 335.89, df = 12, p-value < 2.2e-16Task 4.7
Categorize the variable bmi into an underweight (BMI<18.5), normal weight (18.5≤BMI<25), overweight (25≤BMI<30) and obese (BMI≥30) group. Turn the variable into a factor.
line
Task 4.8
What is the proportion of overweight or obese people according to the categorized BMI? What is the proportion of people ever diagnosed with being overweight (variable ovrwght_ever)? How many overweight people were actually ever diagnosed with being overweight?
bmi_cat <- NULL
bmi_cat[dat$bmi<18.5] <- "Underweight"
bmi_cat[dat$bmi>=18.5&dat$bmi<25] <- "Normal weight"
bmi_cat[dat$bmi>=25&dat$bmi<30] <- "Overweight"
bmi_cat[dat$bmi>=30] <- "Obese"
bmi_cat <- as.factor(bmi_cat)
dat$bmi_cat <- bmi_cat
prop.table(table(dat$ovrwght_ever))
FALSE TRUE
0.6848109 0.3151891 prop.table(table(dat$ovrwght_ever, dat$bmi_cat), margin = 2)
Normal weight Obese Overweight Underweight
FALSE 0.9671907 0.3321033 0.7632275 1.0000000
TRUE 0.0328093 0.6678967 0.2367725 0.0000000Task 4.9
Is there a difference in diabetes prevalence between obese people diagnosed with overweight and those who were never diagnosed? What about self-rated health? How do you explain the results?
(tab <- table(dat$diab_lft[dat$bmi_cat=="Obese"], dat$ovrwght_ever[bmi_cat=="Obese"]))
FALSE TRUE
1 473 694
2 0 7
3 48 253fisher.test(tab)
Fisher's Exact Test for Count Data
data: tab
p-value < 2.2e-16
alternative hypothesis: two.sided(tab <- table(dat$srhgnrl[bmi_cat=="Obese"], dat$ovrwght_ever[bmi_cat=="Obese"]))
FALSE TRUE
1 39 34
2 98 188
3 237 435
4 98 283
5 9 65prop.table(tab, margin=1)
FALSE TRUE
1 0.5342466 0.4657534
2 0.3426573 0.6573427
3 0.3526786 0.6473214
4 0.2572178 0.7427822
5 0.1216216 0.8783784# Chi-square trend test
chisq.test(tab)
Pearson's Chi-squared test
data: tab
X-squared = 39.326, df = 4, p-value = 5.966e-08